Lecture Notes: DNA Computing
DNA Computer Timeline
1994 – Leonard Adleman demonstrates proof-of-concept with first DNA Computer calculation of Travelling Salesman Problem. (His computation chugged along at 100 trillion floating point operations per second)
1997 – Researchers at the University of Rochester develop DNA Logic Gates.
2002 – Researchers at the Weizmann Institute of Science in Rehovot, Israel a programmable molecular computing machine composed of enzymes and DNA molecules.
DNA Computer runs 330 Trillion operations per second
2004 – Researchers at the Weizmann Institute unveil a DNA computer with an input and output module that could theoretically identify cancerous cells and release anti-cancer drugs to counter them.
2009 – Biocomputing systems were combined with silicon based chips for the first time. An enzyme based OR / AND with resets logic system is accomplished with field-effect silicon chips.
DNA Computing: The Silicon computer replacement?
Silicon based computers are rapidly approaching a wall of stagnant growth. Soon, we’ll have encountered the smallest, fastest, microprocessors we can produce. Thus, people have turned to DNA computers.
As it stands right now, DNA computers won’t be replacing the everyday computer anytime soon. While the system is great at storing large amounts of data, there is no reusability of each part. Thus, while transistors in a silicon computer can be reused for multiple calculations, a DNA strand is only used once. Essentially, you have to make a new computer for every new calculation. On the upside, every multi-cellular organism on the planet contains DNA of varying combinations, so we aren’t likely to have a shortage anytime soon.
Additionally, DNA calculations only appear fast because they are massively parallel. In actuality, each reaction is rather slow but because so many solutions can be tried for at once, it takes on the appearance of speed. For a problem that can have hundreds or thousands of possible routes and solutions, each one can be solved simultaneously. Another problem is that DNA computations don’t make it easy to actually see the solution. There is currently no easy way to plug the data from a DNA calculation into an electrical output.
DNA calculations are error prone. Between mismatched pairs and shearing from larger molecules, large scale computations result in a higher chance of errors. The components of the computer are probabilistic and a sub-circuit may give the correct answer only 90% of the time. Additionally, DNA molecules can denature and dissolve into nucleotides. For computations taking an extended amount of time, your computer will be slowly “melting”.
Finally, each stage of DNA computing requires a lot of assistance from a human source. In other words, DNA computers can’t function on their own.
So why DNA?
If DNA computers don’t appear feasible anytime soon, why pursue them? Well, for one thing, the advantages that DNA computers would bring make them worthy enough of a subject.
a) So long as there are multi-cellular organisms, there will be DNA strands to use.
b) DNA bio-chips can be made cleanly, unlike most silicon based computers.
c) DNA computers would be much smaller than modern computers, while holding vast amounts of data.
d) The ability for massive parallel processing is enticing.
e) Low power usage.
As we’ve seen in the computing world, size matters. For the past several decades we’ve gone from colossal transistors to tiny micro computers. A DNA computer would stretch the imagination of “size matters” to the absurd. One pound of DNA material would have the capacity to contain more data than all the electronic computers every built. A DNA computer the size of a teardrop, using DNA logic gates, would theoretically be more powerful than the world’s greatest super computer.
It is anticipated that when a fully functioning DNA computer is released upon the world, it won’t be designed to work with a word processor or basic systems but rather designed to solve extremely complex problems, like that of the travelling salesman.
In terms of power, a DNA based computer uses much less than a standard computer. They DNA hybridization and rebinding is done using stored potential energy within the bonds themselves. You can find a breakdown of estimated power usage on the wikipedia page for DNA Computers.
Just how do DNA Computers Work?
It may be difficult for non-chemist to imagine how DNA can be used to perform computations; therefore examining a few possible implementations of a DNA computer will help in understanding what makes them special.
About DNA
Deoxyribonucleic acid is best known for storing the genetic code of living things. DNA is made of a sequence of nucleotides bound to another sequence of complementing nucleotides in a double helix. Each nucleotide contains one of four bases: adenine (A), cytosine (C), guanine (G), and thymine(T). A and T are complements while G and C are complements. These bases constitute data. As DNA is base 4 and very small (bases are spaced 0.35 nanometers apart), DNA has a data density of about 18 Mbits per inch, or over one million Gbits per square inch if you assume one base per square nanometer. The entire complement of the data is also stored as a form of redundancy.
World's First DNA Computer
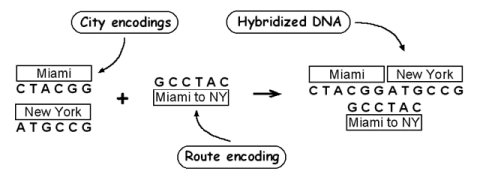
Adleman's first DNA computation solved a traveling salesman problem of seven cities. The method of doing so is surprisingly simple. Each city is represented by a unique sequence of bases. Connections between two cities are created from a combination of the complement of the first half of the sequence of one city, and the complement of the second half of the sequence of a connected city. In this way DNA representing the trip will be created with one strand representing a sequence of cities and the complementing strand representing a series of connections.

The above will create DNA representing all sorts of trips. Some trips may not visit all cities, while other trips may visit some cities more than once. The next step is to process the data until the correct trip remains (one that starts at a certain city and ends in another city after visiting all the others once). The correct answer is obtained by filtering out trips that start and end in the correct cities, then filtering trips with the correct number of cities, and finally filtering out trips that contain each city only once.
DNA Processing with Enzymes
While DNA can be thought of as data, we need to be able to operate on data. One such method is to use enzymes. There are many different enzymes capable of performing many different operations on DNA, such as cutting and pasting. For example, ligase can be used in the above example to allow the strands representing cities and connections to bind together. Continuing from where we left off, the enzyme polymerase can be used to make copies of DNA that begin or end with a certain sequence. This can be used to amplify trips that start and end in the correct cities.
Completing the computation involves sorting the trips by length through gel electrophoresis and picking out trips that visit all cities through affinity purification. These aren't the focus of this paper, but feel free to read more at http://arstechnica.com/reviews/2q00/dna/dna-1.html.
DNA Logic Gates
Adleman’s DNA computer leaves a lot to be desired when compared to silicon computers. A different computation would entail creating a new experiment, or effectively, creating a whole new computer. As a result, much research is being done in creating logic gates out of DNA to form the building blocks of a DNA computer, in much the same way silicon computers are designed. While many different ways of building logic gates have been found, we will just look at one example.
DNAzymes Based Gates
In Adleman’s computation, we looked at how enzymes can be used to modify DNA. Another method is to use DNAzymes, which are also known as DNA enzymes, catalytic DNA, or deoxyribozyme . DNAzymes are DNA that are catalysts and are able to perform operations in much the same way enzymes are. A source of DNA used as input to the gate is able to either promote or inhibit the catalytic action. The action of the catalyst is to cleave some substrate DNA that is present. The substrate is chosen such that once cleaved, it is detectable through fluorescence spectroscopy. This serves as the output.
A NOT gate:
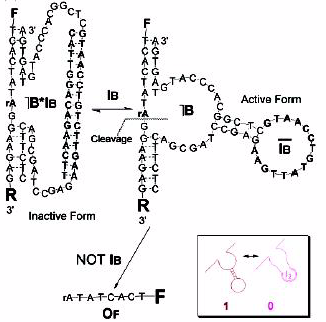
(http://www.ra.informatik.uni-stuttgart.de/~ghermanv/Lehre/Seminar/material/Presentation10/report.pdf)
Links of Interest:
http://computer.howstuffworks.com/dna-computer.htm
http://www.guardian.co.uk/technology/2008/jul/24/computing.research
http://www.nature.com/embor/journal/v4/n1/full/embor719.html
http://www.casi.net/D.BioInformatics1/D.Fall2000ClassPage/DC1/dc.htm
http://publish.uwo.ca/~jadams/dnaapps1.htm
Paper showing feasibility of DNA computers, for the math people:
http://gatekeeper.dec.com/pub/toomany/paper123.pdf